Overview
The main focus of our research is to understand the forces that shape genetic variation across the genome of plants and also how this variation interact to produce adaptive phenotypes. Our work combines computational analysis of high-throughput sequencing data, de novo genome assembly, population and quantitative genetics, and experimental work. We are a highly collaborative lab and work closely together with other labs at SLU in Uppsala where we are physically located, and also with colleagues at other institutes in Sweden, such as Umeå Plant Science Centre, as well as abroad.
De novo Genome sequencing
We are involved in a number of genome projects, including European aspen (Populus tremula), Norway spruce (Picea abies) and timothy (Phleum pratense). Our tasks range from re-sequencing and variant discovery to de novo genome assembly and annotation.
Population and quantitative genomics of adaptation
 Local adaptation plays a fundamental role in maintaining adaptive genetic variation as a response to changing environments but it’s underlying genetic mechanisms remain poorly understood. In this project we aim to identify genetic variation responsible for adaptation to photoperiod in aspen as well as to understand the historical processes that have shaped this variation. We are primarily focusing on phenology traits that are related to the seasonal cycle of growth and dormancy and how these traits varies across a latitudinal gradient across Scandinavia.
Local adaptation plays a fundamental role in maintaining adaptive genetic variation as a response to changing environments but it’s underlying genetic mechanisms remain poorly understood. In this project we aim to identify genetic variation responsible for adaptation to photoperiod in aspen as well as to understand the historical processes that have shaped this variation. We are primarily focusing on phenology traits that are related to the seasonal cycle of growth and dormancy and how these traits varies across a latitudinal gradient across Scandinavia.
A recently started project is aimed at understanding the genetic basis of flowering time variation in two domesticated species of beans, common bean (Phaseolus vulgaris) and scarlet runner bean (Phaseolus coccineus). The two species have their origin in Mesoamerica but have been cultivated in Europe for at least 400 years. During the adaptation to European growing conditions, both species have shifted their flowering time and we are now trying to identify the genetic changes involved in controlling these adaptations.
of flowering time variation in two domesticated species of beans, common bean (Phaseolus vulgaris) and scarlet runner bean (Phaseolus coccineus). The two species have their origin in Mesoamerica but have been cultivated in Europe for at least 400 years. During the adaptation to European growing conditions, both species have shifted their flowering time and we are now trying to identify the genetic changes involved in controlling these adaptations.
 Genomic consequences of linked selection and mating system variation
Genomic consequences of linked selection and mating system variation
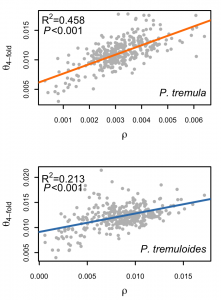
Under the neutral theory, standing genetic variation is the result of a balance between mutation and genetic drift. However, recent studies have show that also natural selection – via positive selection favouring beneficial mutations (genetic hitchhiking) and/or purifying selection against deleterious mutations (background selection) – plays an important role in moulding the landscape of nucleotide polymorphism in many species. We are currently investigating the causes and consequences of linked selection in a number of species of Populus (aspen and poplars) and Picea (spruce) that differ in a number of important characteristics, such as effective population size.
We have also recently initiated a project aimed at understanding how natural selection interacts with mating system variation in shaping the genetic architecture of important adaptive traits. We are investigating these questions in two species of beans, the common bean (Phaseolus vulgaris) and the scarlet runner bean (P. coccineus). Questions of interest in this project include whether selfers experience reduced substitution rates and lower rates of adaptation? Do selfers experience fewer but stronger selective sweeps during adaptation? Do selfiers accumulate higher levels of segregating and fixed deleterious load?
Comparative genomics
The majority of variation in rates of molecular evolution among seed plants remains both unexplored and unexplained. In this project we are analysing a number of large-scale genomics data sets that have recently become available in plants in a phylogenetic framework to understand how and why molecular evolution varies among different plants clades.
 Genomics-assisted breeding
Genomics-assisted breeding
Genomic prediction or genomic selection (GS) is one of the most recent developments in genomics-assisted methods that are aimed at improving breeding efficiency and genetic gains. We have ongoing projects evaliating the utility of genomic prediction for breeding on growth and wood property traits in hybrid Eucalyptus as well as employing related methods to study the genetic architecture of genotype-by-environmnet (GxE) interaction and the genetic phenotypic plasticity in Populus. We are also currently initiating a project aimed at implementing genomic selection through a population-based breeding program in the outcrossing forage grass timothy (Phleum pratense).